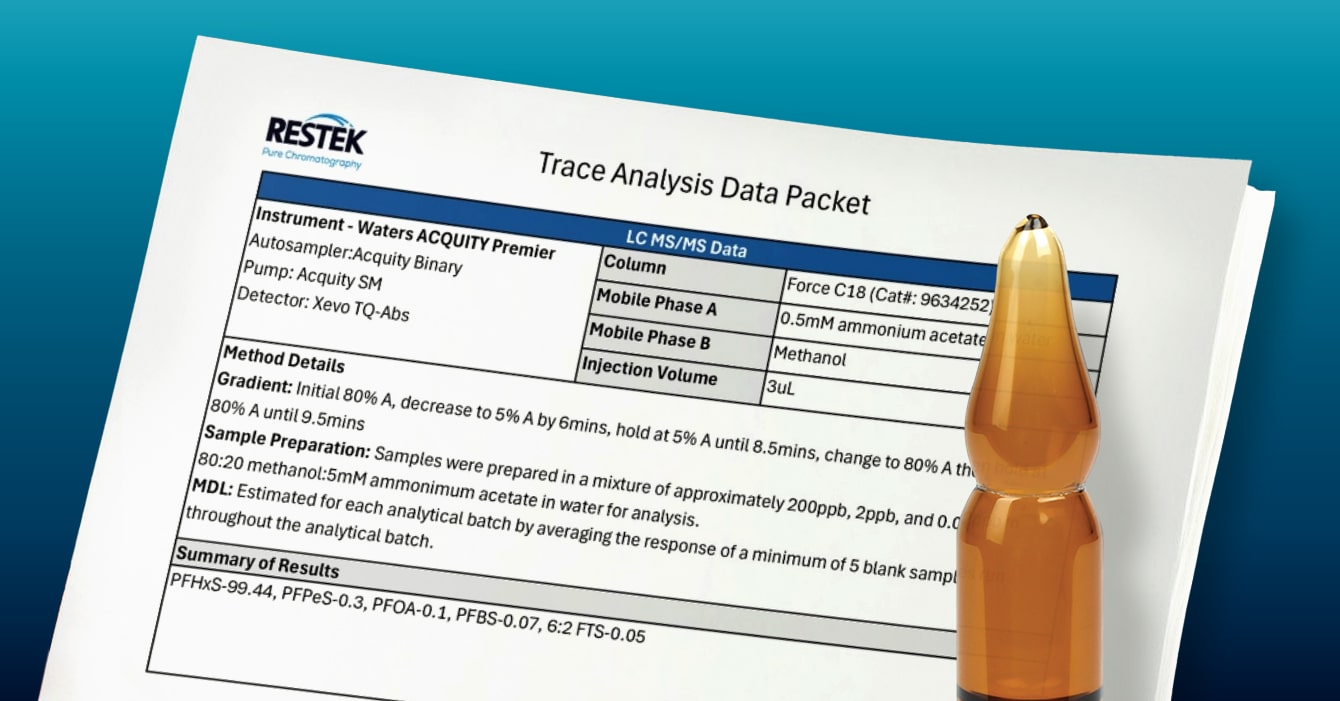
Trace analysis data packets, which are part of our expanded data packs for Restek’s single and multicomponent PFAS reference standards, include detailed information to help you understand the composition of your standards. This article explains how to read your packet, using our br-PFHxS standard (cat.# 30803) as an example.
You can access the trace analysis data packet for your standard by using our documentation search tool and entering your standard’s cat.# and lot#. You can also access your standard’s certificate of analysis (CofA) at the same location.
Restek’s trace analysis data packets include data from two methods, each for a different purpose. The first section of the data pack uses LC-MS/MS for trace analysis of samples. The second section, which is explained later in this article, uses NMR to identify individual isomers.
LC-MS/MS Trace Analysis of Samples
Analytical Method Summary Table
The first table (figure 1) within the trace analysis data packet includes a summary of the analytical method used for trace analysis of samples.
Figure 1: LC MS/MS Data
| LC MS/MS Data | ||
|
Instrument – Waters ACQUITY Premier Autosampler: Acquity Binary Pump: Acquity SM Detector: Xevo TQ-Abs | Column | Force C18 (Cat#: 9634252) |
| Mobile Phase A | 0.5mM ammonium acetate in water | |
| Mobile Phase B | Methanol | |
| Injection Volume | 3uL | |
| Method Details Gradient: Initial 80% A, decrease to 5% A by 6mins, hold at 5% A until 8.5mins, change to 80% A then hold at 80% A until 9.5min. Sample Preparation: Samples were prepared in a mixture of approximately 200ppb, 2ppb, and 0.02ppb in 80:20 methanol:5mM ammonium acetate in water for analysis. MDL: Estimated for each analytical batch by averaging the response of a minimum of 5 blank samples run throughout the analytical batch. | ||
| Summary of Results | ||
|---|---|---|
| PFHxS-99.44, PFPeS-0.3, PFOA-0.1, PFBS-0.07, 6:2 FTS-0.05 | ||
Method detection limit (MDL) is determined in each analytical batch to ensure there is no background contamination within the instrument. This is used to determine the final LOQ.
The Summary of Results for each analyte is measured >LOQ in % wt.
MRM Transitions Table
The second table (figure 2) contains an analyte list and transitions measured by the trace analysis.
Figure 2: MRM Transitions
| Compound | Parent Ion 1 | Daughter Ion 1 | Parent Ion 2 | Daughter Ion 2 | Compound | Parent Ion 1 | Daughter Ion 1 | Parent Ion 2 | Daughter Ion 2 |
| PFBA | 213 | 169 | PFNA | 463 | 219 | 463 | 419 | ||
| PFMPA | 229 | 85 | PFOS | 499 | 80 | 499 | 99 | ||
| 3:3FTCA | 241 | 117 | 241 | 177 | FOSA | 498 | 78 | 498 | 478 |
| PFPeA | 263 | 69 | 263 | 219 | NMeFOSA | 512 | 169 | 512 | 219 |
| PFMBA | 279 | 85 | PFDA | 513 | 219 | 513 | 469 | ||
| HFPO-DA | 285 | 169 | 285 | 185 | NEtFOSA | 526 | 169 | 526 | 219 |
| NFDHA | 295 | 85 | 295 | 201 | 8:2 FTS | 527 | 81 | 527 | 507 |
| PFBS | 299 | 80 | 299 | 99 | 9Cl-PF3ONS | 531 | 351 | 533 | 353 |
| PFHxA | 313 | 119 | 313 | 269 | PFNS | 549 | 80 | 549 | 99 |
| PFEESA | 315 | 83 | 315 | 135 | PFUnA | 563 | 269 | 563 | 519 |
| 4:2 FTS | 327 | 81 | 327 | 307 | NMeFOSAA | 570 | 419 | 570 | 483 |
| 5:3FTCA | 341 | 217 | 341 | 237 | NEtFOSAA | 584 | 419 | 584 | 526 |
| PFPeS | 349 | 80 | 349 | 99 | PFDS | 599 | 80 | 599 | 99 |
| PFHpA | 363 | 169 | 363 | 319 | PFDOA | 613 | 319 | 613 | 569 |
| ADONA | 377 | 85 | 377 | 251 | NMeFOSE | 616 | 59 | ||
| PFHxS | 399 | 80 | 399 | 99 | NEtFOSE | 630 | 59 | ||
| PFOA | 413 | 169 | 413 | 369 | 11Cl-PF3OUdS | 631 | 451 | 633 | 453 |
| 6:2 FTS | 427 | 81 | 427 | 407 | PFTrDA | 663 | 169 | 663 | 619 |
| 7:3 FTCA | 441 | 317 | 441 | 337 | PFDoS | 699 | 80 | 699 | 99 |
| PFHpS | 449 | 80 | 449 | 99 | PFTeDA | 713 | 169 | 713 | 669 |
Quantification is based on response from transition corresponding to Parent 1 > Daughter 1. Secondary transition is used for qualitative identification.
Trace Analysis Summary
The third table (figure 3) is summary of the trace analysis. We have color coded the columns to help explain the table. The columns will be uncolored in your data packet.
Figure 3: Trace Analysis Summary
| Reporting Limit (%wt) | LOQ (ng/g) | Measured Conc (ng/mL) | Dilution factor | % Composition | |
| PFHxS | 0.05 | 0.01 | 2.40 | 100 | 99.45 |
| PFPeS | 0.05 | 0.01 | 0.73 | 1 | 0.30 |
| PFOA | 0.10 | 0.01 | 0.25 | 1 | 0.10 |
| PFBS | 0.05 | 0.01 | 0.17 | 1 | 0.07 |
| 6:2 FTS | 0.05 | 0.05 | 0.14 | 1 | 0.06 |
| PFBA | 0.10 | 0.01 | 0.04 | 1 | <0.1 |
| PFMPA | 0.10 | 0.05 | N.D. | 1 | <0.1 |
| 3:3FTCA | 0.10 | 0.03 | N.D. | 1 | <0.1 |
| PFPeA | 0.10 | 0.06 | N.D. | 1 | <0.1 |
| PFMBA | 0.10 | 0.03 | N.D. | 1 | <0.1 |
| HFPO-DA | 0.10 | 0.03 | N.D. | 1 | <0.1 |
| NFDHA | 0.10 | 0.05 | N.D. | 1 | <0.1 |
| PFHxA | 0.10 | 0.20 | N.D. | 1 | <0.1 |
| PFEESA | 0.10 | 0.01 | N.D. | 1 | <0.1 |
| 4:2 FTS | 0.10 | 0.02 | N.D. | 1 | <0.1 |
| 5:3FTCA | 0.05 | 0.05 | N.D. | 1 | <0.05 |
| PFHpA | 0.10 | 0.31 | N.D. | 1 | <0.1 |
| ADONA | 0.10 | 0.01 | N.D. | 1 | <0.1 |
| 7:3 FTCA | 0.10 | 0.05 | <LOQ | 1 | <0.1 |
| PFHpS | 0.10 | 0.31 | N.D. | 1 | <0.1 |
| PFNA | 0.05 | 0.01 | N.D. | 1 | <0.05 |
| PFOS | 0.05 | 0.06 | N.D. | 1 | <0.05 |
| FOSA | 0.13 | 0.02 | N.D. | 1 | <0.125 |
| NMeFOSA | 0.13 | 0.01 | N.D. | 1 | <0.125 |
| PFDA | 0.10 | 0.01 | N.D. | 1 | <0.1 |
| NEtFOSA | 0.10 | 0.01 | N.D. | 1 | <0.1 |
| 8:2 FTS | 0.05 | 0.05 | N.D. | 1 | <0.05 |
| 9Cl-PF3ONS | 0.05 | 0.05 | <LOQ | 1 | <0.05 |
| PFNS | 0.05 | 0.01 | <LOQ | 1 | <0.05 |
| PFUnA | 0.10 | 0.01 | N.D. | 1 | <0.1 |
| NMeFOSAA | 0.13 | 0.01 | N.D. | 1 | <0.125 |
| NEtFOSAA | 0.13 | 0.01 | N.D. | 1 | <0.125 |
| PFDS | 0.05 | 0.01 | N.D. | 1 | <0.05 |
| PFDOA | 0.10 | 0.03 | N.D. | 1 | <0.1 |
| NMeFOSE | 0.13 | 0.09 | N.D. | 1 | <0.125 |
| NEtFOSE | 0.13 | 0.08 | <LOQ | 1 | <0.125 |
| 11Cl-PF3OUdS | 0.05 | 0.05 | N.D. | 1 | <0.05 |
| PFTrDA | 0.10 | 0.38 | N.D. | 1 | <0.1 |
| PFDoS | 0.05 | 0.01 | N.D. | 1 | <0.05 |
| PFTeDA | 0.10 | 0.025 | N.D. | 1 | <0.1 |
Reporting Limit (red column): Typically, 0.05% for sulfonates, 0.10% carboxylic acids, and 0.125% for sulfonamides.
LOQ (blue column): Limit of Quantitation (LOQ) of the instrument. Value is equal to the lowest calibrator or the batch specific MDL, whichever is greater.
Measured Conc (green column): Measured concentration on instrument. <LOQ means the peak was detected but below the analyte’s LOQ. Not Detected (N.D.) analyte was below the detection limit of the instrument.
Dilution factor (orange column): The dilution factor each analyte was quantified from. Target analyte typically had to be diluted to ensure the measured concentration was within the linear range of the analyte.
% Composition (purple column): Results in % wt for each target analyte. Values are calculated from Measured Concentration * Dilution Factor. Final values are normalized to 100%. Reporting limits are applied after normalization.
Overlay MRM Chromatogram
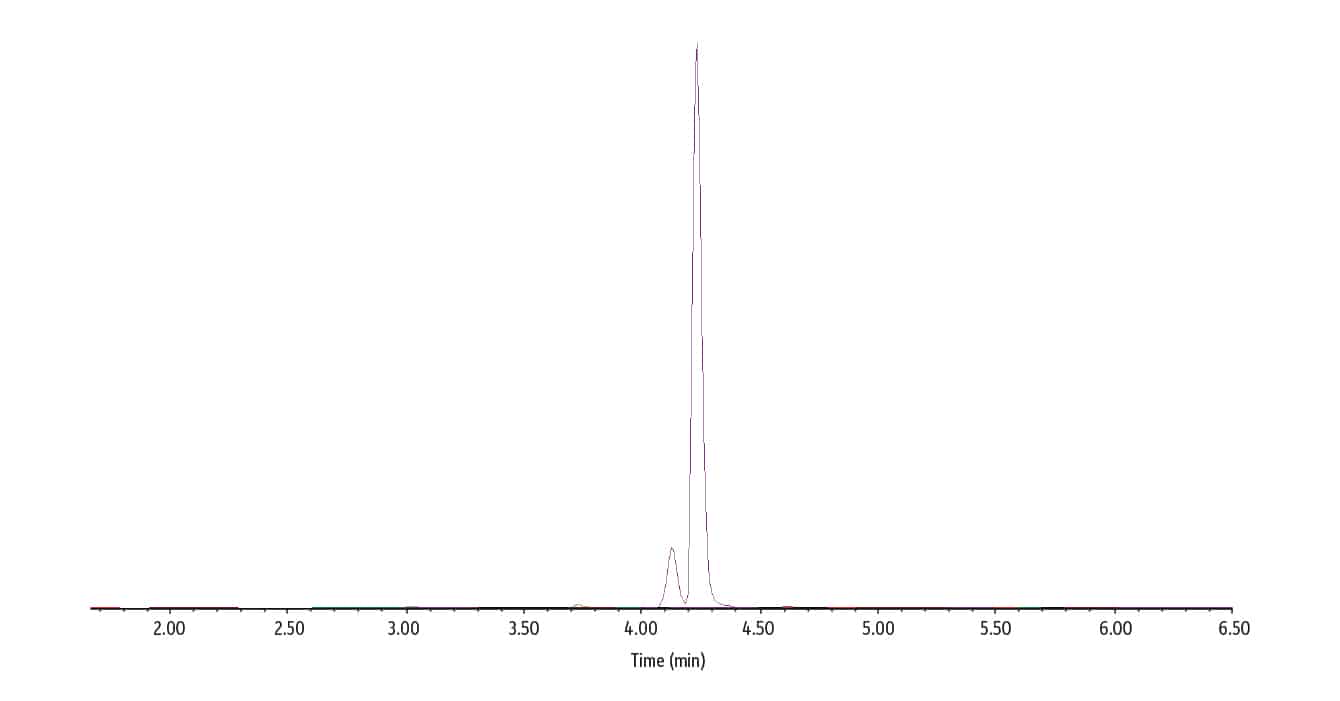
The next data (figure 4) is an overlay MRM chromatogram of the trace analysis. The full scan data can be found in the CofA. The data is typically from the most concentrated value measured.
Figure 4: Overlay MRM Chromatogram
NMR Analysis to Identify Individual Isomers
NMR Table
Figure 5: NMR Table
The fourth table (figure 5) includes NMR data, which lists sample preparation information and summary of results.
| NMR |
| Isomeric ratios determined by NMR will differ from ratios determined chromographically due to unresolved separation. Identification of individual isomers is based on published work. (doi:10.1016/j.chemosphere.2007.06.096) Samples were prepared at approximately 10mM with an approximate MDL of 1%wt. |
| Summary of Results |
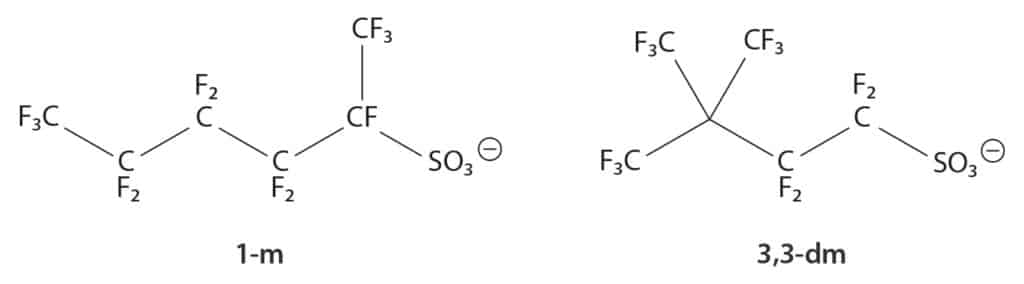
| The following isomers were identified: 1-m, 2-m, 3-m, 4-m, and 3,3-dm. |
In the Summary of Results, the number denotes distance from end-functional group. m stands for “methyl” and dm for “dimethyl,” indicating the isomer contains either a single-branching CF3 group or two CF3 groups respectively.
Figure 6: Example Numbering Convention for PFAS Isomers
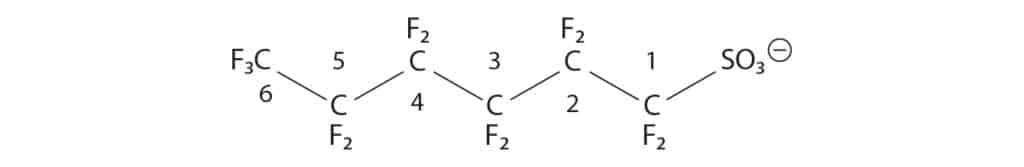
Figure 7: Example Isomer Naming Convention
NMR Data
The final data (figure 8) included in the trace analysis data packet is additional NMR data.
Figure 8: NMR Data-Annotated
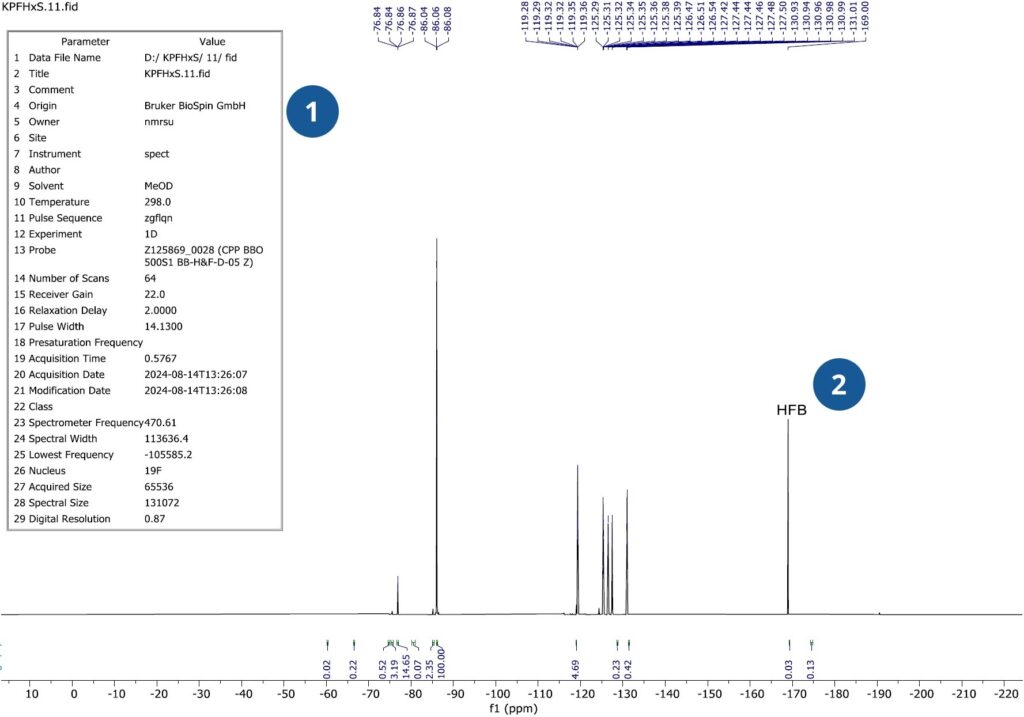
Figure annotations:
- Acquisition parameters for NMR data.
- Hexafluorobenzene (HFB) added as an internal standard to determine relative chemical shifts.
Full Support for Your Restek PFAS Standards
For more information about our expanded data packs and PFAS reference standards, review our full reference standard FAQ. Additionally, if you have questions about your specific standard, contact our Technical Service team.



